Electric Steam Boiler / Generator
Click Here for Steam Generator Models
.
Please take me to the Airtorch® Models Page or Take me to the Steam Generator Models Page.
Or use a hybrid. Contact MHI.
If a large kW boiler is required only for heating, one may consider changing to process heat generators with or without moisture to save energy and water. Please view these pages for high-volume steam generators like the OAB-50 or OAB-100.
High-Efficiency Electric Steam Boiler / Generator – One Atmosphere Boiler for heating kettles and vats. Use hot air/gas or steam. MHI manufactures all types of steam generators.
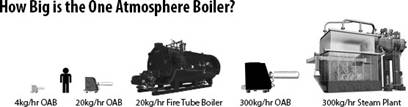
Compare Efficiency. Three types of efficiencies are essential to consider. (1) the efficiency of the steam generation device where the GHGA, OAB®, and MightySteam® excel; (2) the efficiency of the downstream process that the steam is used for; and (3) the overall water usage efficiency and propensity to recycle the water. The one-atmosphere steam-gas generator/boiler OAB® performs well in all three categories. Better conversion factors allow OAB® units to use fewer resources to produce more steam. A standard OAB steam output at 350°C is almost at the theoretical efficiency of power conversion for OAB steam generation! The OAB’s rapid startup (generally within a minute from a cold) also considerably reduces energy usage.
 |
It was getting to temperature quickly and faster, resulting in lower activewear and related productivity with higher kinetics. The output velocity of the 550°C OAB steam is very high – almost 40m/s or more. This feature is used for uniform kettles/heat exchangers’ uniform heating and reaching large piping distances without significant temperature loss. The heating rate for industrial vats and kettles significantly affects the overall process economics. A general rule of thumb for productivity from traditional pressure boilers is – (0.35 Gallons of a heated product like water)/kW-hr. For the OAB, it is closer to – (1 Gallon/kW-hr), i.e., almost three times (numbers may be lower or higher as they are situationally dependent and one needs to evaluate on a case-by-case basis). Traditionally an 80-200 kW boiler is used for vat-heating by microbreweries and juice producers. Imagine cutting down to 1/3rd the power required. This is a new 2021 year technology.
Note that 1 Kilowatt (KW)= 3,414 BTU/hour. At the lower end of the clean burn, about 0.4 Kg of CO2 is produced for every 1 kWh of combustion. For a 100 KW combustion boiler running continuously and burning fuel for 24 hrs., ~ 960 Kg (close to a ton) of CO2 may be produced per hour. Home-gas-burners burn at aboHome gas burners natural gas is used. If home-gas-fired burners are used for ~2 hrs., then about 2-4 Kg or more of CO2 could be made in that time. Please note that modern burners are restricted on the amount of CO made, but venting is essential whenever a burner is used. Please review, for example, published links – https://www.abe.iastate.edu/extension-and-outreach/carbon-monoxide-poisoning-gas-fired-kitchen-ranges-aen-205/.
How much does energy cost? The price of energy depends on the source but is not that different between different forms of commonly used energy types that one can use (e.g., gas or electric). Comparisons are given, for example, here. In the US, when comparing gas and electricity, a rule of thumb is that gas energy is about 75% of the cost compared to the same quantity of electric energy. However, electric machines OAB® or even electric boilers are generally much more efficient than gas heaters. Additionally, electric heaters can be efficient/accurately controlled. For natural gas heating, about 117 pounds of CO2 are produced for one million BTU (293 Kwh) energy produced by the combustion of n. The OAB® produces no CO2. A unit called Boiler-Power(BP) is used in the boiler world. (1BP ~ 10kW). So 8 BP-rated boiler is approximately an 80 kW boiler. https://mhi-inc.com/oab-instant-high-temperature-steam-models/
https://mhi-inc.com/oab-instant-high-temperature-steam-models/
Example of energy savings: Microbrewery or Cottage Cheese Milk owners want to use much less energy and maintain an ecologically low carbon footprint. A typical microbrewery may use kettles and vats heated with steam (or high-quality hot air). Sometimes steam heating can replace high-quality Air Heating for the same kW’s. Airtorch units are less expensive than the corresponding steam units for the same kW.
General instructions for Beer. Unless steam is required, a hot air device may be considered easy to use and afford savings by avoiding water use. A typical microbrewery may use kettles and vats heated with steam (or high-quality electric Airtorch® hot air) with the following sequence. Note that the scenario below is optimized for the lowest power-use possibility. MHI offers one to several hundred kg/hr steam generators or Airtorch® Units.
Step 1: Heat approximately 600 gallons of water from 75°F to 170°F in a kettle in 6 hours ~ requires 24 kW for a well-insulated kettle (i.e., one OAB -36, MVTA-36 or two OAB-12 or two MVTA925-12 units for use overnight). Lower power equals a longer heat-up time if the insulation is adequate. 12 kW = 40968 BTU/hr. If you are a specialized microbrewery making 100 gallons daily, each vat may require only two f heating time.
Step 2: Add grain to 600 gallons & maintain 150° F-170°F temperature in kettle 1 hr ~ requires 12 kW (one OAB-12 or MVTA925-12) to maintain the temperature at 170°F. This depends on how much cold grain is added. Please check with your Brewmaster for accuracy. Grain can be preheated independently with an efficient hot air device or steam before VAT heating.
Step 3: Remove grain and bring the liquid to a mild boil for 6 hours (from 170°F to 210°F) – requires 12kW depending on how much steam is made during the spot (one OAB-12 or MVTA925-12). The immediate introduction is not feasible for all brews or with air to avoid combustion.
Using a more efficient steam generator like the OAB® allows one to reduce operating costs while demonstrating ecological sensitivity. With an OAB® or Airtorch, please remember it is an on-off device (on demand), unlike many traditional boilers. Remember also that it is your choice to use the medium of heating, steam, or air. Contact MHI.
Note* Powers stated are approximate. How much savings per kW saved? This page may be used to calculate the energy and power required.
When using steam, i.e., an OAB®, please remember it is an on-off device (on demand), unlike a traditional boiler which often has to be idled. Save enormous electric and water usage by using this on-demand feature. Use only when required.
How does one increase the rate of production?
Short Answer: Increase the temperature of the process gas or steam. Yes, this can be done with the OAB steam or an Airtorch®.
Detailed Answer: Please click on the Steam Calculator page.
HGA and OAB Steam Generator Models. A list of applications for the best steam available.
 MightySteam® Products (Industrial and R&D Use)
MightySteam® Products (Industrial and R&D Use)
Steam boilers and generators utilizing 20th-century technology are innately inefficient. Combustion processes are typically inefficient or polluting. . Even with economizers and the best designs, most fire tube and water tube boilers can only approach 85% efficiency. Low-efficiency ratings, large footprints, long startup times, and safety concerns have been the norm for any application requiring steam. Steam boilers generate steam through the use of combustion and high pressure. MHI’s full line of patented steam boilers and generators generate pure, superheated steam at 1-atmosphere pressure without any discharge.
- Electric Steam Generation – MHI’s Steam Boiler and Generator lines generate steam without combustion. Or use the Airtorch®. No NOX by-products here.
- Pure Clean Steam – The highest quality, low moisture content dry steam available is offered by many of MHI’s steam generators.
- High Temperatures – Steam temperatures are available from 300°C up to 1300°C. Scalable units provide steam without ever-increasing operating pressure!
- 21st Century Efficiency – MHI’s steam products are 99%+ outlet to output efficient. Convert electricity to steam while minimizing losses resulting from combustion.
- Excellent Design – Most MHI electric boilers are small enough to be near the demand source. Save money by not having to route steam piping.
Electric steam generators are now a reality. With small device footprints and unparalleled feature sets, products such as our OAB can revolutionize processes and reduce inefficiencies in transporting steam through pipes, idling boilers, and the combustion process. With outputs available to over 200kg/hr of 1300°C steam, MHI’s One Atmosphere Boilers can fulfill many small to moderate steam demands.
For more information, see Superheated Steam or One Atmosphere Electric Steam Boiler. HGA and OAB Steam Generator Models